In this section, we will see how to use py3Dmol to visualize MD trajectories, either as an animation, as well as using a slider to observe each individual step. We first need to import the required modules:
import py3Dmol as p3d
import ipywidgetsAssuming that we have the MD trajectory saved in a PDB file, we need to write a routine which reads all the steps from the PDB file and returns them as text, as required by the Py3Dmol viewer.
def read_pdb_file(file_name):
"""Reads the MD trajectory from a PDB file."""
pdb_file = open(file_name, "r")
data = pdb_file.read()
pdb_file.close()
return dataWe can now create an animation of the dynamics.
pdb_file_name = '../../data/md/bithiophene.pdb'
pdb_data = read_pdb_file(pdb_file_name)
viewer = p3d.view(width=600, height=300)
viewer.addModelsAsFrames(pdb_data)
viewer.animate({"loop": "forward"})
viewer.setViewStyle({"style": "outline", "width": 0.05})
viewer.setStyle({"stick":{},"sphere": {"scale":0.25}})
viewer.show()If instead we would like to observe each step (in this case, each step is 1 ps) individually, we need a routine which reads one configuration at a time from the PDB file.
def read_pdb_index(file_name, index=0):
"""Reads one configuration defined by its index from a PDB file."""
pdb_file = open(file_name, "r")
current_index = 0
data = ""
for line in pdb_file:
if index == current_index:
data += line
if 'ENDMDL' in line:
if index == current_index:
break
current_index += 1
pdb_file.close()
return dataAdditionally, we need a routine which creates a py3Dmol viewer for each configuration.
def return_viewer(file_name, step=0):
pdb_data_i = read_pdb_index(file_name, step)
viewer = p3d.view(width=600, height=300)
viewer.addModel(pdb_data_i)
viewer.setViewStyle({"style": "outline", "width": 0.05})
viewer.setStyle({"stick":{},"sphere": {"scale":0.25}})
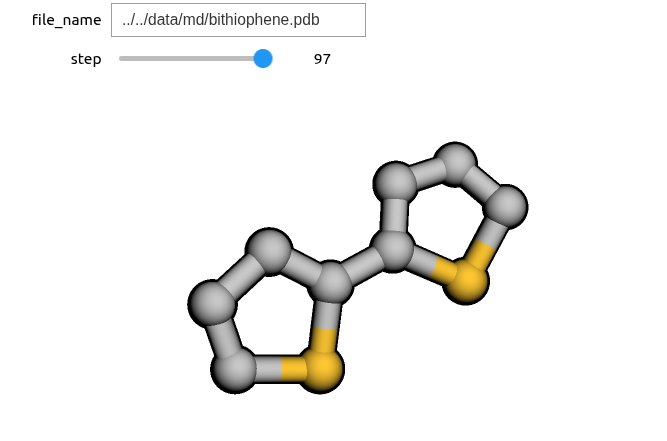
viewer.show()Now we can create a widget with a slider that allows us to visualize one configuration at a time. To do so, we make use of the ipywidgets module which allows to interactively change the parameters of a Python routine. In our case, we use ipywidgets to allow the user to set the trajectory file name and step index required by the return_viewer routine which generates the py3Dmol viewer. The step is set using a slider widget which allows the user to select any value between the defined limits (note that the slider only works in a Jupyter notebook).
total_steps = 100 # number of configurations in the trajectory file
ipywidgets.interact(return_viewer, file_name=pdb_file_name,
step=ipywidgets.IntSlider(min=0, max=total_steps, step=1, value=49))